Binan lab
Investigating spatial and temporal mechanisms of gene regulatory networks.
We develop technologies to interrogate gene expression regulation in single cells in their spatial context. We ask how the local environment is responsible for the activation of different genetic pathways and we look for the consequences of these transcriptional events on cellular differentiation and migration.
We use precision gene editing tools such as Cas9 and Cas12 in high throughput screens. We develop imaging tools to collect new data in a spatial context, and the computational tools to analyze these new data types. We develop our methods in the contexts of cancer and neurodegenerative diseases.
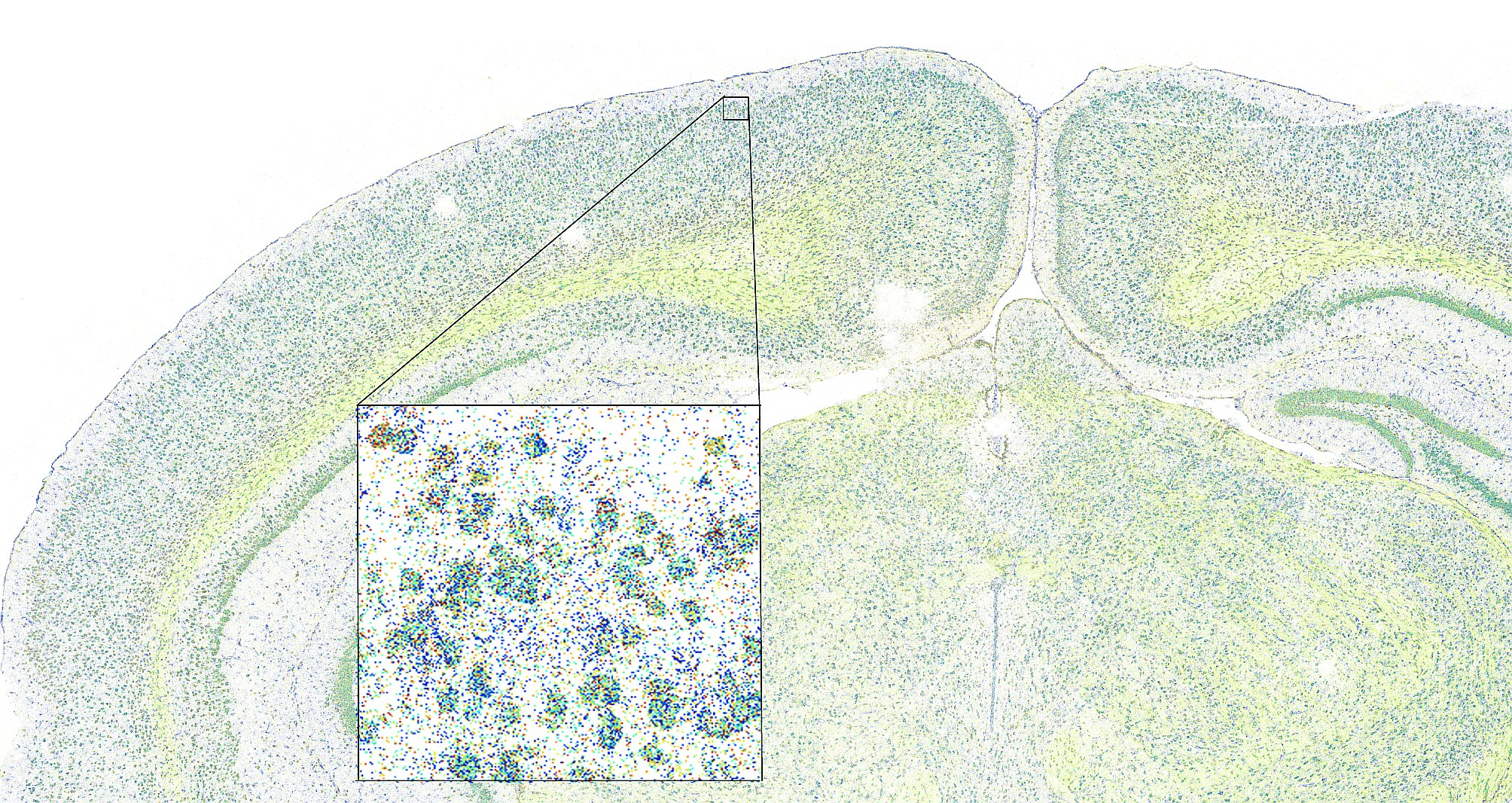

Understanding the mechanisms of metastasis initiation with spatial transcriptomics and genetic perturbations.
The majority of cells at the primary site of a tumor have an epithelial phenotypes, but many cells undergo partial, transient transitions to mesenchymal-like phenotypes. While epithelial, tumor cells proliferate fast and are well adhered to their neighbors, but when they become mesenchymal, proliferation slows down and cells start migrating. In this state, they can reach new sites, where they revert to epithelial traits and restart proliferating, forming a metastatic lesion. This process is highly dynamic, and strongly influenced by the microenvironment. We are investigating the genetic mechanisms that allow cells to reversibly transition between epithelial and mesenchymal phenotypes.
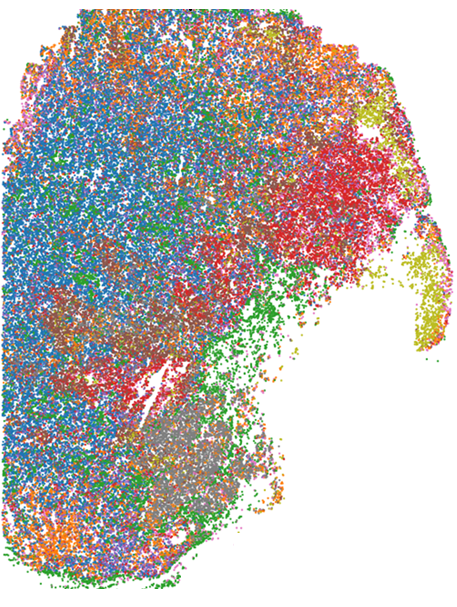

Many genes can be expressed as a variety of isoforms that contain different fragments, or exons, of the sequence of the same gene. Beyond differences in total number of molecules of RNA from a given gene, the function of alternatively spliced genes strongly depends on differences in their sequence. We develop technologies to understand the regulation and effect of alternative splicing in the context of neurodegenerative diseases.
Spatial technologies are designed to allow interrogating the influence of cellular neighborhoods on genetic pathways and resulting phenotypes. This analysis is limited to a snapshot of the spatial environment of the sample at the moment it was collected, and the information about past interactions of cells with their environment is lost. We develop tools to investigate how past cell-cell interactions shape present transcriptional activity in cells during development.